Highlight achievements: 3)Structure and function of membrane proteins and protein complexes----Institute of Biophysics Chinese Academy of Sciences

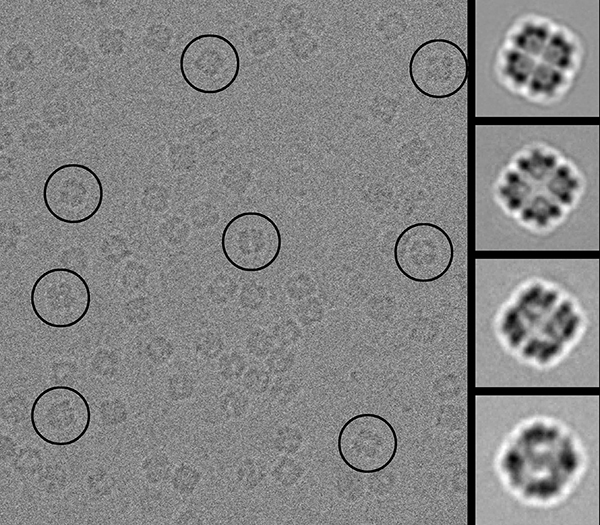
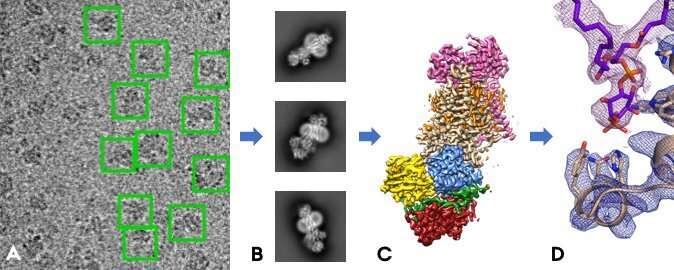
Structure Determination (A) Cryo-EM image of wt Hsp26 complexes. In the... | Download Scientific Diagram

Protein structure by optimized negative staining (OpNS) EM. Electron... | Download Scientific Diagram

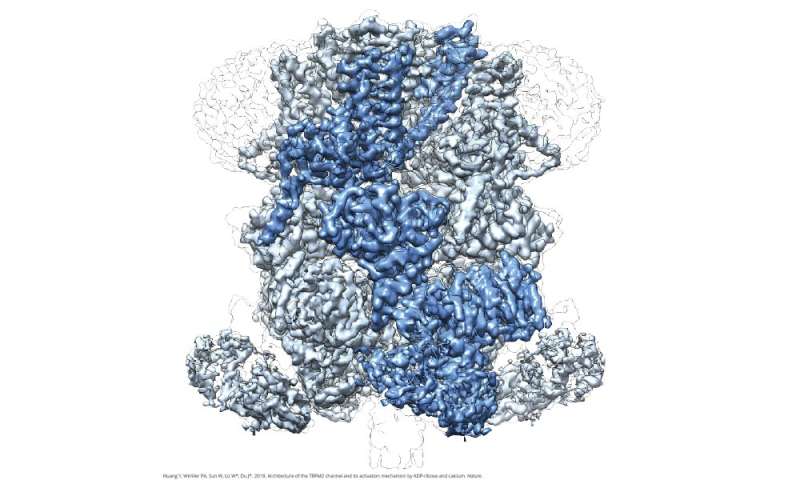
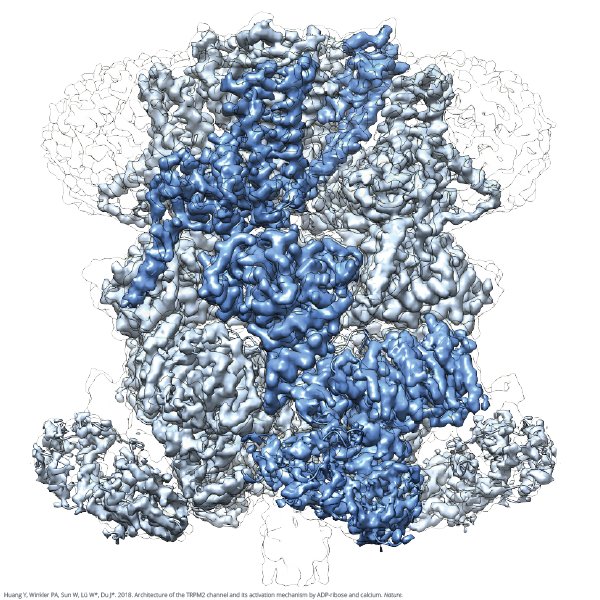
A New Player Emerges in Mapping Protein Structures, Thanks to Berkeley Lab Technology | Berkeley Lab
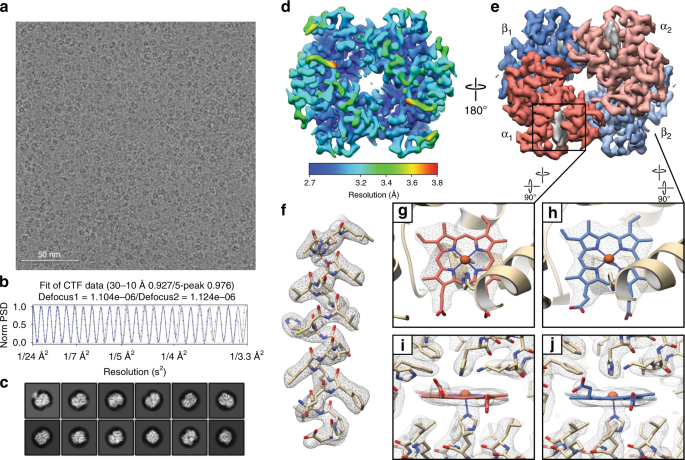
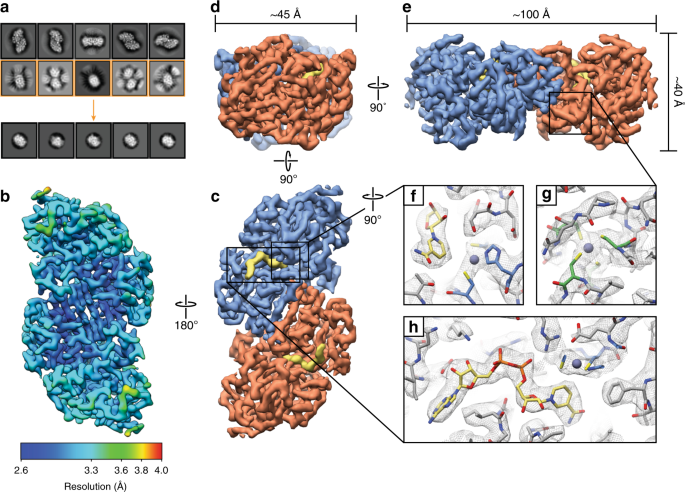
High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM | Nature Communications
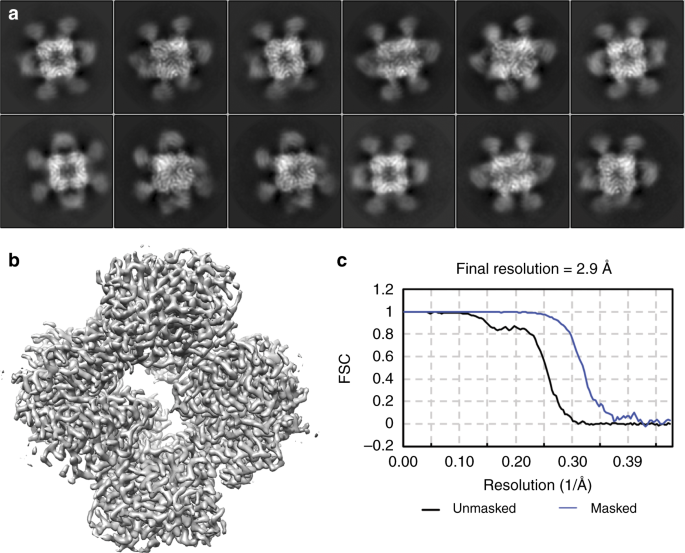
A 3.8 Å resolution cryo-EM structure of a small protein bound to an imaging scaffold | Nature Communications